Latest Findings of NKU's Research Team in the Field of Autophagy Published in Leading Journal The Plant Cell

Prof. Li Lei from the College of Life Sciences, Nankai University formed an international joint research team with Prof. Harvey Millar of the University of Western Australia and Prof. Marisa Otegui of the University of Wisconsin. Using stable isotope metabolic labeling and a protein turnover analysis platform created by mass spectrometry technology, in conjunction with cell biology, metabolomics analysis and biochemical experimental evidence, the team systematically compared the differences in the protein turnover of model plant Arabidopsis thaliana between wild-type and autophagic mutants, and discovered a new autophagic degradable protein substrate. The research paper entitled "Defects in autophagy lead to selective in vivo changes in turnover of cytosolic and organelle proteins in Arabidopsis" was recently published online in The Plant Cell, an international prestigious academic journal.
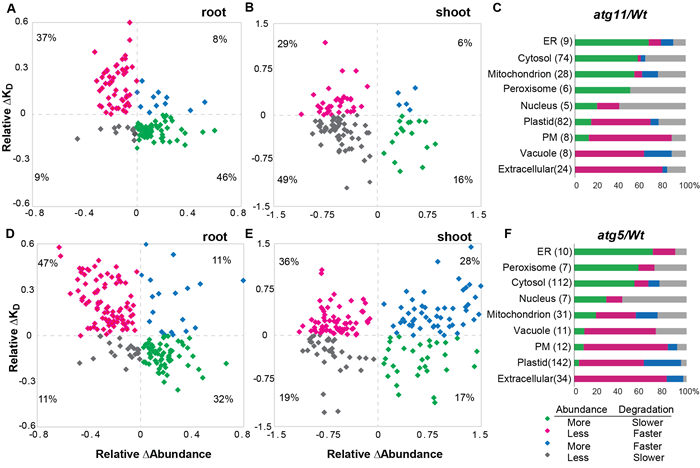
Figure 1. Conjoint analysis of protein abundance and turnover of autophagic mutants.
Prof. Li Lei's research team systematically compared and analyzed the protein abundance and turnover rates of the wild-type and autophagic mutants atg5 and atg11 of the model plant Arabidopsis thaliana through the protein turnover analysis platform, and found that the degradation rate of less than half of the upregulated protein was reduced (Figure 1). The collaborative analysis of protein abundance and degradation rate showed that 122 varieties of proteins accumulated and downregulated in autophagic mutants, including previously confirmed autophagic substrate ribosomes, proteasomes, mitochondria, and other organelles. It is noteworthy that the study discovered new autophagic degradation protein substrates that had not previously been reported in plant cells, including a variety of proteases during glycolysis, chaperones, and other proteins (Figure 2). This laid the foundation for further in-depth study of the recognition mechanism of autophagic degradation.
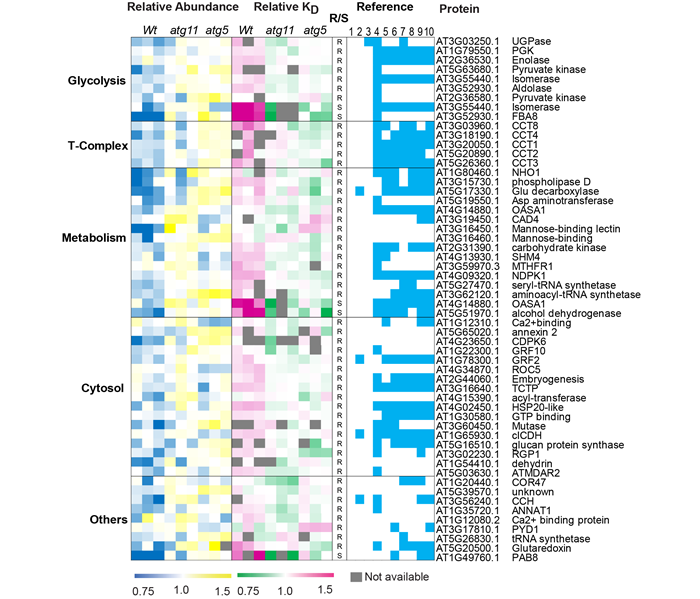
Figure 2. Discovery of new autophagic degradation substrate proteins.
The study validated the newly discovered autophagic substrate glycolytic enzyme FBA8 through cell biology and biochemical autophagic substrate identification strategies. It was found that fluorescently labeled FBA8 can enter autophagosomes and autophagic vesicles, while inhibitors for vacuole acidification can effectively downregulate their autophagic degradation (Figure 3). Furthermore, through the comparison of the functionally inactivated mutants of autophagic core protein ATG5 and ATG11, it was found that ATG11 has a more significant effect on regulating protein degradation of mitochondria and chloroplasts, and that BFA globule-like special membrane structures were found in ATG5 functional mutants. It showed that ATG5 and ATG11 made possible the in-depth research on the working mechanism of autophagy degradation and specific regulation of other vesicle transport pathways within cells.
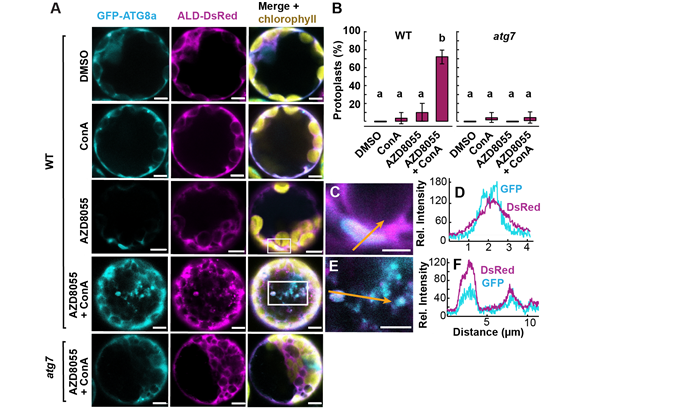
Figure 3. Validation of new autophagic degradation substrate proteins.
Prof. Li Lei is the first and co-corresponding author of the paper. Qin Yu and Chen Zhirui , graduate students of the research team participated in the research work.
Link to the paper: https://academic.oup.com/plcell/advance-article-abstract/doi/10.1093/plcell/koac185/6619566
(Edited and translated by Nankai News Team)









